Structure of YF1. a Structure of YF1 in its dark-adapted state as

Download scientific diagram | Structure of YF1. a Structure of YF1 in its dark-adapted state as resolved by X-ray crystallography 13. The location of the different domains, of the flavin mononucleotide (FMN), of the cofactor adenosine diphosphate (ADP), and of the phosphoaccepting histidine 161 are indicated. b Light induced conformational changes of the LOV photosensor domain refined from X-ray solution scattering 22. The changes are maximal at the C-termini that feed into the Jα helices (dashed arrows). The coloring is according to the root mean square deviation of the alpha carbons from publication: Sequential conformational transitions and α-helical supercoiling regulate a sensor histidine kinase | Sensor histidine kinases are central to sensing in bacteria and in plants. They usually contain sensor, linker, and kinase modules and the structure of many of these components is known. However, it is unclear how the kinase module is structurally regulated. Here, we use | Secondary Protein Structure, Bacterial Proteins and Protein Conformation | ResearchGate, the professional network for scientists.
4,4'-Dihydroxydiphenylmethane-d10
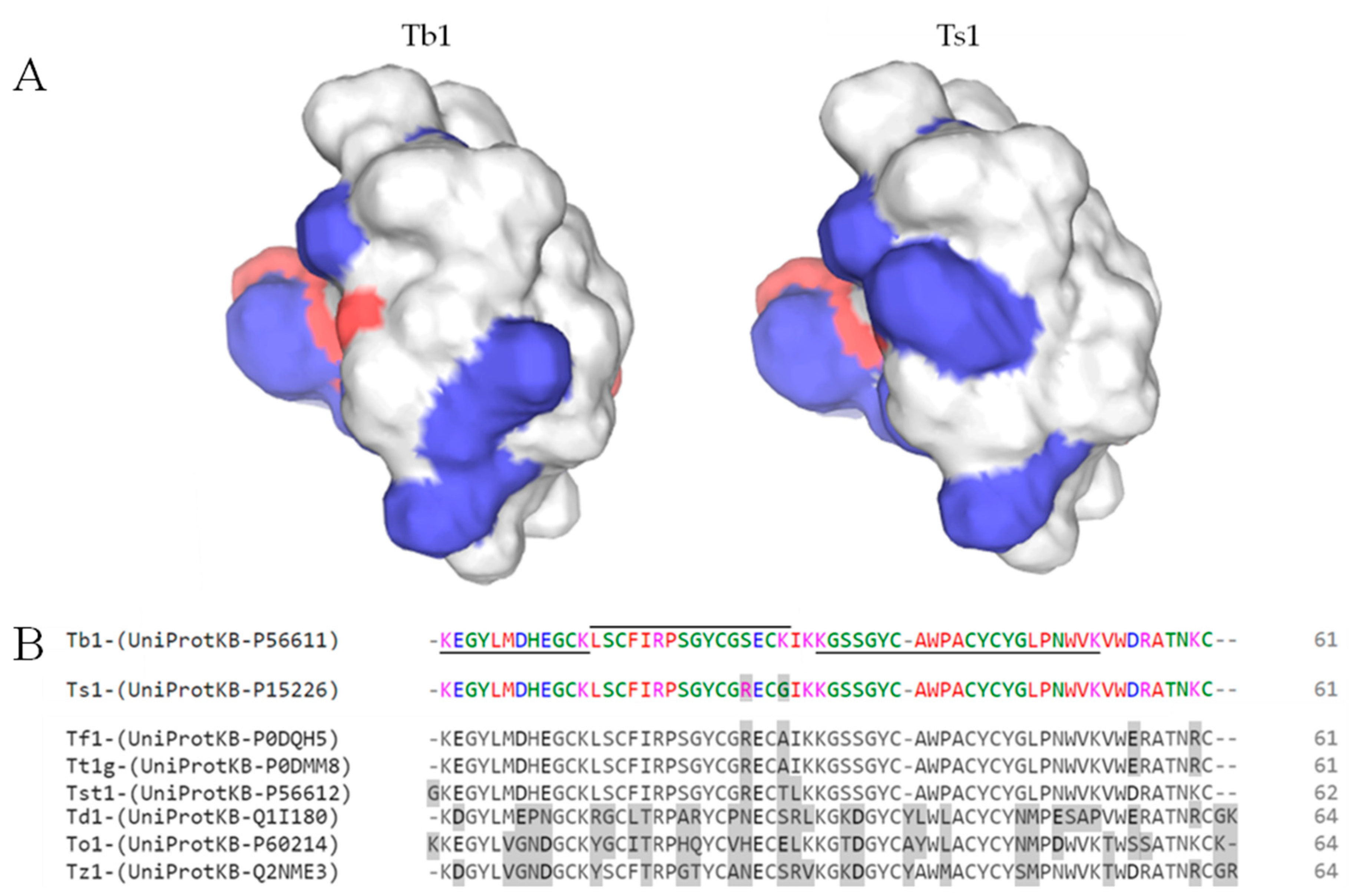
Toxins, Free Full-Text
Molecules, Free Full-Text
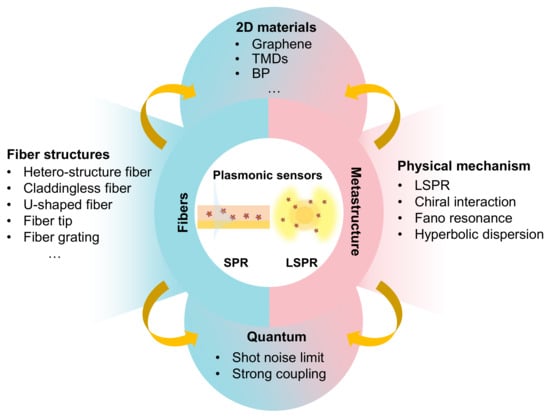
Sensors, Free Full-Text

Structure of the ATP synthase catalytic complex (F1) from

Structure of photoreceptors. (A) The LOV2 domain of Avena sativa
IJMS, Free Full-Text

Staircase solutions and stability in vertically confined salt-finger convection, Journal of Fluid Mechanics

Structure of photoreceptors. (A) The LOV2 domain of Avena sativa

Signal transduction in light–oxygen–voltage receptors lacking the adduct-forming cysteine residue

Architecture and function of the pREDusk and pREDawn expression

Structural photoactivation of YF1. Blue light causes sequential

Structure of the ATP synthase catalytic complex (F1) from Escherichia coli in an autoinhibited conformation

Matthijs PANMAN, Scientist, PhD, VSL - Dutch Metrology Institute, Delft, VSL, Temperature and Humidity









